This is the most exciting step in your analysis of DNA sequencing data: genome assembly!
Before building the genome of your samples, first check metrics such as coverage, read depth, N50 and L50 to have a sense of where your analysis may be headed. Then, pick your assembly strategy, depending on your samples and goals. Finally, annotate your assembled genome, or call its variants.
It’s a lot of fun!
REMINDER: This glossary is divided in three parts, covering the three major computational steps in the analysis of DNA sequencing data:
3. ASSEMBLING A GENOME
17 – SEQUENCING COVERAGE: the percentage of the genome that was sequenced at least once.
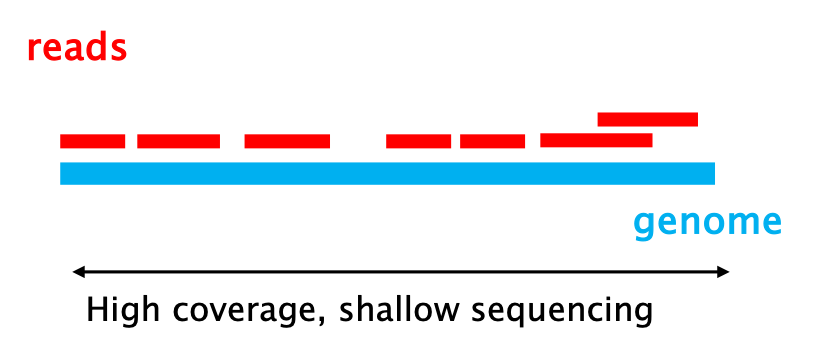
18 – SEQUENCING READ DEPTH: how many times a specific base has been sequenced, that is how many reads contain that base.
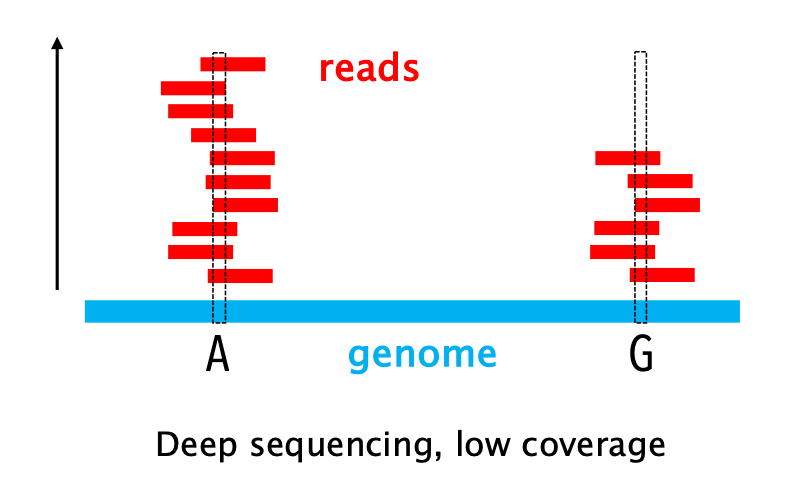
NOTE: sequencing coverage and read depth are often used interchangeably. Although related, the two metrics are not synonymous, as I recently explained.
19 – N50: the length of the shortest contig (or scaffold) such that all contigs of that length or longer sum to at least 50% of the total assembly size..
20 – L50: the number of contigs (or scaffolds) to reach N50.
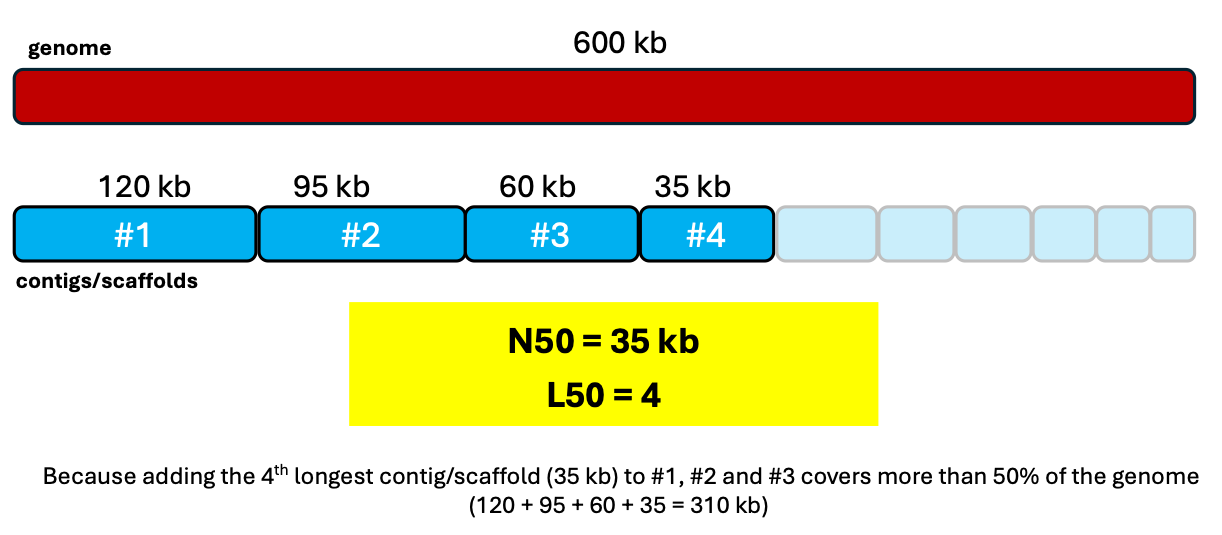
NOTE: N50 & L50 are metrics of how contiguous or fragmented a genome assembly is. Higher N50 and lower L50 are generally better.
21 – GENOME ASSEMBLY: Reconstructing a genome by stitching back together reads using their overlapping sequences. Two main approaches exist: reference-based and de novo assembly (along with other less common hybrid strategies).
22 – REFERENCE-BASED ASSEMBLY: the assembly is guided by a reference sequence (described in the “READING THE GENOME” episode), to which the reads are mapped to determine their order and orientation. This approach reduces the computational burden, making the process much faster. However, adhering to a reference may hide unexpected structural variations in the genome sequenced, especially in repetitive or highly divergent regions.
23 – DE NOVO ASSEMBLY: the order and orientation of overlapping reads are determined without a reference, that is without a priori knowledge of the genome under scrutiny. This approach is the forced choice when no reference is available, as for giant genomes, or to tease out structural variations between the sequenced genome and the reference. While powerful, it is more computationally intensive (therefore slower) and often struggles to reconstruct large repetitive regions. It generates contigs and scaffolds.
24 – CONTIG: a continuous sequence built by stitching together overlapping reads. A contig (from “contiguous”) contains no gaps or unknown bases. While small genomes (virus, bacteria) can sometimes be represented as a single contig, sequencing of larger genomes typically produces many contigs that are later assembled into scaffolds.
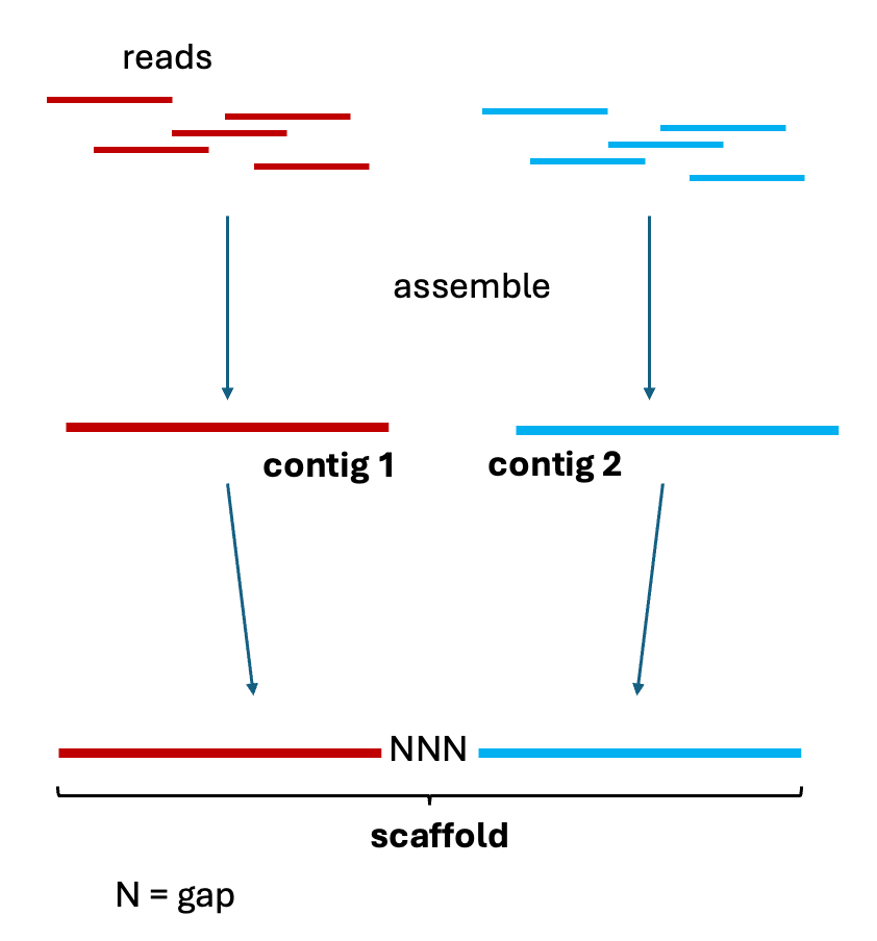
25 – SCAFFOLD: a chain of contigs in the appropriate orientation and order but interrupted by gaps of unknown sequence (typically indicating that coverage is not complete). These gaps must be resolved by sequencing targeted to the boundaries of these regions or by using other complementary techniques.
26 – GENOME ANNOTATION: the process of identifying and describing the functional elements of a de novo assembled genome, such as protein-coding genes, non-coding genes (here’s an overview of ncRNAs, in progress) and regulatory elements,
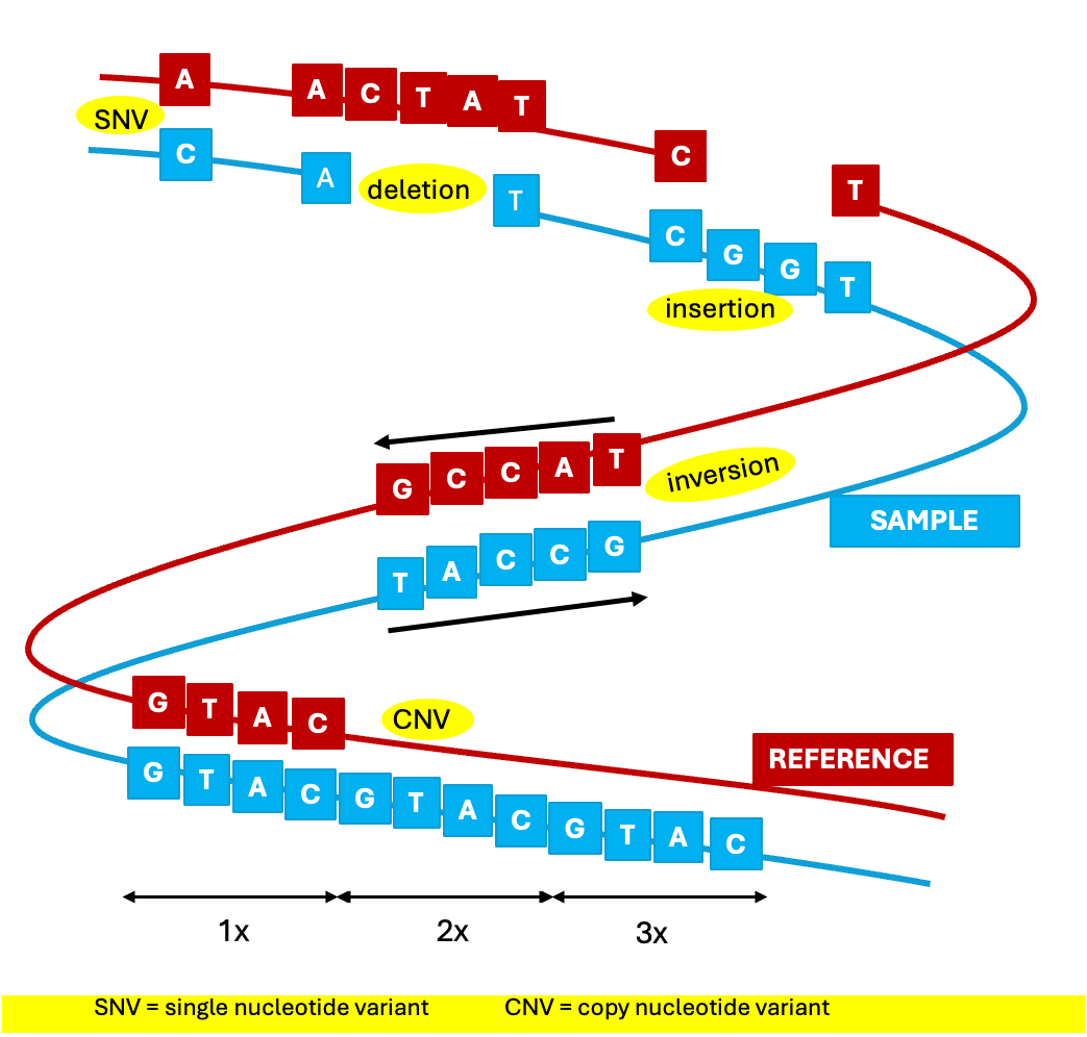
27 – VARIANT CALLING: the computational process that identifies the differences between a reference genome and the genome of a sample. A high sequencing read depth is crucial for the variant to be called with confidence. Variants called range from single or multiple nucleotide variant (SNV and MNV) to deletion, inversions, inversions and other rearrangements.
WHAT NOW
I hope this short three‑episode glossary on reading, mapping, and assembling a genome helps you as you take your first steps in this fascinating world… and maybe nudge you towards a new career path!
If you want to learn more about the story of DNA sequencing, have a look at another series I am working on: A Chronicle of DNA sequencing in 5 anniversaries!


